Research
See also: CV (pdf).
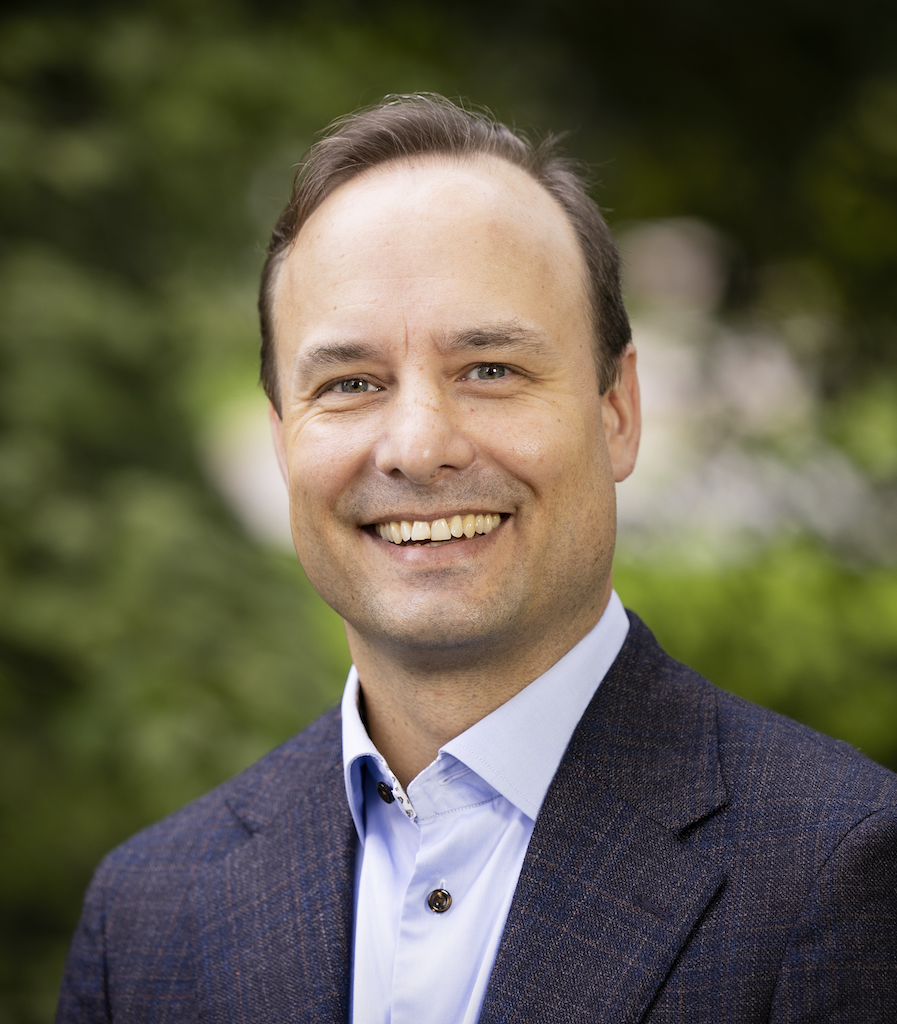
Research
I am particularly interested in Scientific Computing in the intersection with Data-driven research and Data Science. I have extensive experiences in many aspects of Scientific Computing in general, in Numerical Modeling and -Analys in particular, as well as to some extent in High-Performance Computing. My main focus of applications are in the Biosciences at broad, but I’ve also taken an interest in traditional computational Engineering applications.
Current active research projects include Bayesian approaches for compute intensive data-driven models in epidemics, including in particular prediction, and multiscale modeling and parameterization of living cells, where spatial stochasticity is an important aspect of the modeling.
I am currently the main supervisor for 3 PhD-students:
- Erik Blom (started 2021). Eriks project is Scalable computational modeling of living cells.
- Gesina Menz (started 2022). Gesinas project is Data-driven modeling of living cells.
- Vaishnavi Divya Shridar (started 2024). Divyas project is Towards model-based analysis in wastewater epidemiology: the specifics of antimicrobial resistance.
I was the main advisor of
- Robin Marin who successfully defended his thesis Computational Modeling, Parameterization, and Evaluation of the Spread of Diseases (2022).
- Jing Liu who successfully defended her thesis Towards Fast and Robust Algorithms in Flash X-ray single-particle Imaging (2020). This project was run joint with Janos Hajdu’s group at the Department of Cell and Molecular Biology.
- Pavol Bauer who successfully defended his thesis Parallelism in Event-Based Computations with Applications in Biology (2017).
I am also the secondary advisor for Anna Frigge, Alfred Andersson, and Helena Andersson. I was previously the secondary advisor of
- Adrien Coulier. PhD thesis Multiscale Modeling in Systems Biology: Methods and Perspectives (2021).
- Afshin Zafari. PhD thesis Advances in Task-Based Parallel Programming for Distributed Memory Architectures (2018).
- Stefan Widgren. PhD thesis Studies on verotoxigenic Escherichia coli O157 in Swedish cattle. From sampling to disease spread modelling (2016).
- Lina Meinecke. PhD thesis Stochastic Simulation of Multiscale Reaction-Diffusion Models via First Exit Times (2016).
- Marcus Holm. Licentitate thesis Scientific computing on hybrid architectures (2013).
On the national side I was previously an elected member of the Young Academy of Sweden for the period 2016–2021. See my alumni presentation web-page there.
MSc/BSc-theses:
- Approximate Bayesian Computation for Data-Driven Epidemiological Models by Christoph Nötzli (2023, MSc Data Science)
- Investigating the Estimation of the infection rate and the fraction of infections leading to death in epidemiological simulation by Jakob Gölén (2023, MSc Engineering Physics)
- Cell-sorting in grid-based time-continuous cell population models by Joel Olofsson (2022, MSc Engineering Physics)
- Computational modelling of quorum sensing using cascade delay by Nils Axelsson and David Mårsäter (2022, BSc Engineering Physics)
- Tumörspridning med artificiell evolution: Warburgeffekten och cancercellers metabolism by David Näsström and Marcus Medhage (2022, BSc Engineering Physics)
- Towards Hybrid Modeling of Avascular Tumours by Erik Blom (2021, MSc Computational Science)
- Implementing multithreading for a fast multipole method using OpenMP by Ludwig Ridderstolpe (2021, BSc Computer Science)
- Performance of Adaptive Fast Multipole Method in three dimensions for time-dependent problems by Zain Nawas (2021, MSc Computational Science)
- Comparing Priority Queues with support for priority updates at arbitrary indexes by Erik Granberg (2021, BSc Computer Science)
- Heterogeneous Multiscale Method in Markovian event-based models – With applications in tumor modeling by An Khang Bui (2020, MSc Numerische Mechanik, Technical University of Munich)
- A parallel implementation of spatially distributed stochastic chemical kinetics by Pontus Melin (2020, BSc Computer Science)
- Practical complexity of the Fibonacci heap in a simulation and modelling framework by Elwira Johansson (2020, BSc Computer Science)
- Bayesian inference in Epidemics: consistency and convergence by Samuel Bronstein (2019, MSc (eq.), Applied Mathematics, ENS Paris)
- Bayesian Parametrisation of In Silico Tumour Models by Jonas Radvilas Umaras (2018, MSc Computational Science).
- Computational modeling of avascular tumours using a hybrid on-lattice framework for cell-population dynamics by Lina Viklund (2018, MSc Engineering Physics).
- Mathematical modeling of interactions between colonic crypts by Martin Edin and Nils Erlanson (2017, BSc Engineering Physics)
- Multiscale Stochastic Neuron Modeling: with applications in deep brain stimulation by Aleksandar Senek (2017, MSc Engineering Physics)
- Bayesian Parameterization in the spread of Diseases by Robin Eriksson (2017, MSc Engineering Physics)
- Computational Stochastic Morphogenesis by Yakup Saygun (2015, MSc Engineering Physics)
- High Performance Computing aspects of Single Particle Machine Learning by Marcus Näslund (2015, MSc Computational Science)
- Efficient Parameter Inference for Stochastic Chemical Kinetics by Debdas Paul (2014, MSc Theoretical Biological Physics/Computational Systems Biology)
- Towards mesoscopic modeling of firing neurons: a feasibility study by Emil Berwald (2014, MSc Engineering Physics)
- GPU-Parallel simulation of rigid fibers in Stokes flow by Ronny Eriksson (2014, BSc Computer Science)
- Parallelization and performance in simulation of disease spread by animal transfer by Fredrik Pasanen and Magnus Söderling (2012, BSc Computer Science)
Previously
I became an Associate Professor in 2014, being previously promoted to Docent in 2013 in Scientific computing. I originally joined UPMARC in 2011 with the aim at bringing problems from Scientific Computing into a form suitable to modern multicore/manycore computers, and vice versa, to develop and analyze algorithms and techniques suitable to such cards with interesting applications in mind. Research outputs here include, amongst others,machine learning methods in imaging with X-ray lasers, auto-tuning in CPU/GPU implementations of adaptive fast multipole methods, and shared memory approaches for event-based algorithms.
The Linnaeus center of excellence UPMARC
⇒ Focus area Application Performance
⇒ ⇒ Project group Parallel Algorithms
Before that, I was a PostDoc at the the Linné FLOW Centre where I started in September 2009 to work on computational modeling of multiphase flow for two immiscible fluids and a surface active agent. For example, this would be the correct model when considering a mixture of oil/water and a detergent.
Before that I was also briefly involved in Anna-Karin Tornberg’s project concerning simulating fibers suspended in fluids.
As a graduate student I studied methods for computing numerical solutions to stochastic descriptions of chemical reactions. The underlying mathematical description is a continuous-time Markov chain and the equation governing the probability density is called the Master Equation. Unfortunately, the master equation cannot be solved numerically for more than, say, five molecular species due to the exponential growth of work and memory requirements (‘curse of dimensionality’). Stochastic descriptions of chemical reactions are needed to describe the chemical processes taking place inside living cells with few copy numbers of each molecular species. Usual models for cell simulation are based on the reaction rate equations which form a system of nonlinear ordinary differential equations. Such models ignore the stochastic fluctuations in the cells and are therefore less accurate.
Computational systems biology group.
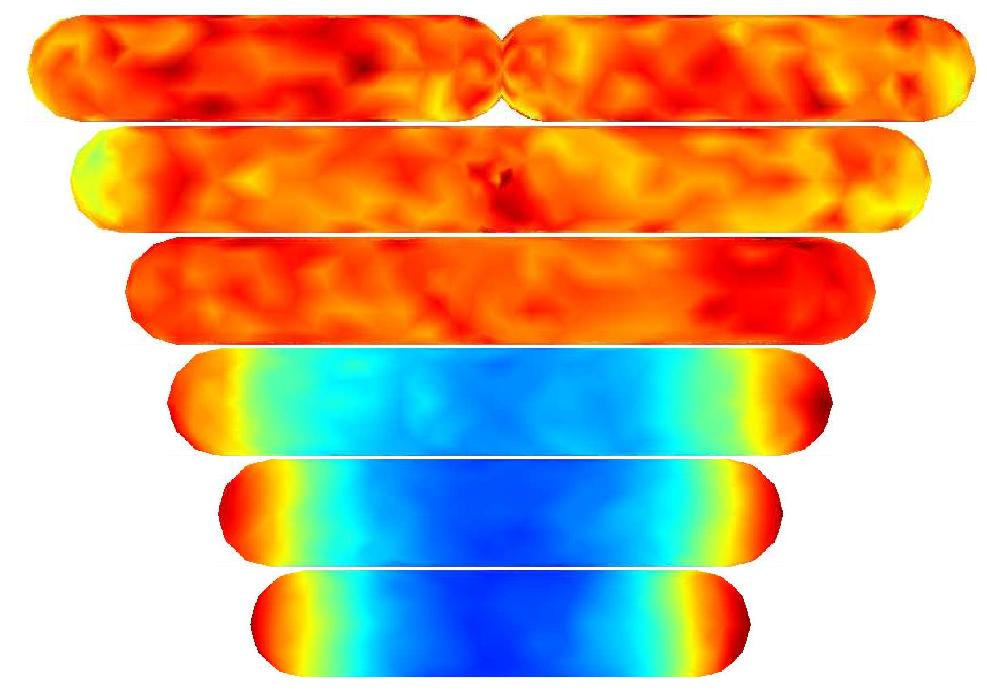
I was also involved in a project joint with the Division for Electricity and Lightning Research, Uppsala University called “Electric power generation from winds”. The ultimate goal of the project is to provide more efficient wind turbines. I have been working together with Paul Deglaire and, lately, Anders Goude. The work has resulted in a two-dimensional random vortex method simulating fluid flows around general airfoils at a quite high Reynolds number. My contribution has been focused around a user-friendly and very efficient implementation of the fast multipole method.
